PATENTS
1. L Cui, K Schleyer, J Liu, Heparanase compounds and methods of use. US provisional patent 2022.
2. L Cui, J. Liu, and K.A. Schleyer. Probes for heparanase. US provisional patent. 2019.
3. L. Cui, J. Liu, Y. Wang, and P. Deenik, Near Infrared small molecule probes for the detection of cellular senescence. US provisional patent. 2017.
PEER REVIEWED PUBLICATIONS
Noninvasive NIR Imaging of Senescence via In Situ Labeling, Jun Liu, Xiaowei Ma, Chao Cui, Zixin Chen, Ying Wang, Philip R Deenik, Lina Cui, Journal of Medicinal Chemistry, (2021).
Noninvasive NIR Imaging of Senescence via In Situ Labeling, Jun Liu, Xiaowei Ma, Chao Cui, Zixin Chen, Ying Wang, Philip R Deenik, Lina Cui, Journal of Medicinal Chemistry, (2021).
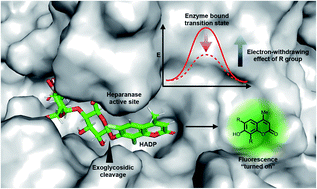
Ultrasensitive small molecule fluorogenic probe for human heparanase, Jun Liu, Kelton A. Schleyer, Tyrel L. Bryan, Changjian Xie, Gustavo Seabra, Yongmei Xu, Arjun Kafle, Chao Cui, Ying Wang, Kunlun Yin, Benjamin Fetrow, Paul K. P. Henderson, Peter Z. Fatland, Jian Liu, Chenglong Li, Hua Guo and Lina Cui, Chem. Sci, 2020, Advance Article
[18F]-SuPAR: A Radiofluorinated Probe for Noninvasive Imaging of DNA Damage-Dependent Poly(ADP-ribose) Polymerase Activity
Adam J. Shuhendler, Lina Cui, Zixin Chen, Bin Shen, Min Chen, Michelle L. James, Timothy H. Witney, Magdalena Bazalova-Carter, Sanjiv S. Gambhir, Frederick T. Chin, Edward E. Graves, and Jianghong Rao*,Bioconjugate Chem. 2019, 30, 5, 1331–1342
From alpha to beta: identification of amino acids required for the N-acetyllactosamine-specific lectin-like activity of bundlin, Romney M. Humphries, Michael S. Donnenberg, Jonathan Strecker, Elena Kitova, John S. Klassen, Lina Cui, Thomas P. Griener, George L. Mulvey and Glen D. Armstrong*, Molecular Microbiology, 2009. 72(4): p. 859-868.



















![41598_2019_38511_Fig1_HTML[1].png](https://static.wixstatic.com/media/7b382e_5987f29029be482db81e9b5c03284e94~mv2.png/v1/crop/x_369,y_0,w_503,h_503/fill/w_165,h_150,al_c,q_85,usm_0.66_1.00_0.01,enc_avif,quality_auto/41598_2019_38511_Fig1_HTML%5B1%5D.png)
![bc-2019-000896_0001[1].gif](https://static.wixstatic.com/media/7b382e_c47e2766241f4edb9b72fb88b72d11ff~mv2.gif)
![cjc-2016-0417f1[1].jpg](https://static.wixstatic.com/media/7b382e_094323e7abcc456c8a0ea42d4599393e~mv2.jpg/v1/crop/x_188,y_0,w_1525,h_1525/fill/w_165,h_150,al_c,q_80,usm_0.66_1.00_0.01,enc_avif,quality_auto/cjc-2016-0417f1%5B1%5D.jpg)





![300992_1_En_5_Fig1_HTML[1].gif](https://static.wixstatic.com/media/7b382e_965263303cb34c81bc249932d9463a41~mv2.gif)